Introduction
Introduction
BioHackathon Germany is an annual event that brings together life scientists, bioinformaticians and data managers / data stewards from Germany and around the world. Organised by de.NBI & ELIXIR-DE, it offers an intense week of hacking with over 150 participants (on-site and online) working on diverse and exciting projects. The aim is to produce results that address challenges in bioinformatics and life science research. Scientists from companies are also encouraged to participate and submit project proposals connecting their work to open science activities and initiatives.
Scope
BioHackathon Germany’s activities are driven by dynamic sessions, where life scientists, bioinformaticians and data managers / data stewards meet, discuss and implement ideas and projects during intensive and productive hacking sessions. To be selected, projects need to demonstrate alignment with the research, infrastructure, training or RDM topics offered by de.NBI & ELIXIR Germany and its service centres.
Goals
-
Promote the development of an open source data integration infrastructure that will accelerate scientific innovation. This includes areas such as FAIR, identifiers, metadata standards, ontologies and metadata catalogues.
-
Engage technical people from the bioinformatics community within and beyond de.NBI / ELIXIR-DE to collaborate on topics of common interest aligned with de.NBI / ELIXIR-DE activities.
-
Strengthen and enable interactions and collaborations with ELIXIR, NFDI and other bioinformatics related projects through joint activities.
-
Provide networking opportunities for all participating attendees
Enhancing FAIR (Meta-)Data Practices in Life Science by Improving RO-Crates Support in Federated Storage Systems
The growing complexity and volume of life science research data present major challenges to managing, sharing, and reusing data across modern storage infrastructures. While enabling collaborative research, data sovereignty and institutional control over datasets have become increasingly important. Moreover, adherence to the FAIR Principles (findable, accessible, interoperable, and reusable) has become a necessity for modern data storage. However, many current storage systems, including federated infrastructures, lack comprehensive support for standardized metadata descriptions. This creates barriers to effective data discovery and cross-platform interoperability.
Research Object Crates (RO-Crates) are a lightweight, community-driven method of packaging research data alongside rich, structured metadata using JSON-LD and Schema.org vocabularies. Currently, most RO-Crate implementations focus on the attached format, in which the metadata and data are packaged together. Others are built only to parse the JSON-LD metadata itself. The detached format has limited support because not all datasets are present at the current location, which complicates data access and validation. Additionally, existing tools lack sophisticated pagination mechanisms when dealing with many files and do not emphasize ingestion workflows for detached formats. They also struggle with scalability, large dataset handling, and distributed metadata management when deployed in federated storage environments, limiting their practical application in modern research infrastructures. Thus, improved library support is necessary to address large-scale, distributed research scenarios.
These limitations will be mitigated by improving the RO-Crate tooling to better support federated storage architectures and ingestion processes. These processes will be able to automatically extract metadata and populate search indices across distributed storage nodes. The improved tooling will support attached and detached RO-Crate formats, as well as paginated RO-Crates. It will enable large-scale research objects that exceed traditional packaging limits. This approach enables researchers to maintain comprehensive metadata descriptions while accommodating the practical constraints of distributed storage systems and network transfer limitations in federated environments. The project will establish guidelines for handling very large datasets within RO-Crate frameworks and address scalability challenges specific to life science data, which may include genomic sequences, high-resolution imaging datasets, proteomics data, and longitudinal experimental collections. Implementation will target federated data storage solutions, such as the Aruna platform, which is used by various NFDI consortia and will serve as the primary testbed for validating enhanced RO-Crate workflows within distributed storage environments.
This project directly supports the strategic vision of ELIXIR-DE and de.NBI by advancing interoperable bioinformatics infrastructures that facilitate cross-border data sharing and collaborative research within the European life sciences community. The enhanced RO-Crate implementation aligns with ELIXIR’s commitment to developing standardized data management practices and metadata frameworks that enable seamless integration across distributed computational resources. By focusing on federated storage architectures and FAIR data principles, the initiative contributes to de.NBI’s mission of providing sustainable, scalable bioinformatics services that can accommodate the growing data volumes characteristic of modern life science research. Furthermore, the project’s emphasis on interoperable metadata management and distributed data ecosystems directly supports the European Open Science Cloud (EOSC) objectives of creating a federated, cross-disciplinary research infrastructure. The implementation of paginated RO-Crates and improved ingestion workflows will enhance the technical foundation necessary for EOSC’s vision of seamless data discovery and access across European research institutions, while maintaining the data sovereignty and institutional control requirements essential for sensitive biological datasets.
Ideas / Discussions
During our participation in the Biohackathon Germany 2025, our working group engaged in extensive discussions focused on the practical and technical challenges of integrating RO-Crates in federated storage systems for life science data. The following sections present the central questions and ideas that emerged from these collaborative sessions. They reflect the real-world needs and obstacles faced by researchers and developers striving to advance FAIR data practices in distributed environments, and highlight areas where current standards and tooling may require further development.
Handling RO-Crates with Extremely Large Datasets and Integrated Metadata
A recurring topic in our discussions was the management of RO-Crates that encapsulate extremely large datasets, particularly when the metadata file itself grows to an impractical size that becomes difficult to process and maintain, leading the group to identify the need for some form of pagination or segmentation.
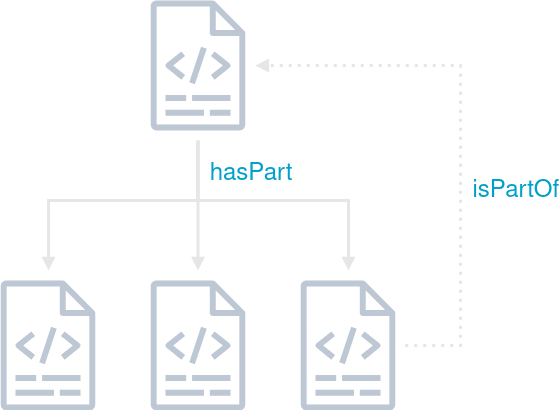
Fig. 1: Schematic representation of a nested RO-Crate including a backwards pointing reference to the parent RO-Crate.
While the RO-Crate specification provides mechanisms for creating and referencing subcrates, a significant challenge arises in how these subcrates can reference their parent crates. Typically, subcrates are unaware of the existence of the top-level crate, which complicates hierarchical organization. One proposed solution was to utilize the isPartOf property to establish a link to the parent RO-Crate, although this property only allows referencing by @id. Addressing these limitations is essential for scalable metadata management in federated systems, and further work is needed to develop robust strategies for handling large, interconnected RO-Crate structures.
Integrating Content Consistency Checks in File Metadata
Another important question was how to ensure content consistency and integrity within the file metadata of RO-Crates. The group discussed the desirability of including properties with each data entity that could be used to validate the integrity of the data entity’s content. While creating a custom “data integrity profile” referenced in the @context section was considered, it was deemed too time-consuming to be prioritized as a high-priority task for the event. Instead, a more pragmatic approach was suggested by directly referencing established terms such as Checksum, ChecksumAlgorithm, and checksumValue from the SPDX vocabulary in the @context property of the RO-Crate, and applying these to each data entity.
Example:
[...]
"@context": [
"https://w3id.org/ro/crate/1.2/context",
{
"checksumAlgorithm": "https://spdx.org/rdf/terms/#checksumAlgorithm",
"checksumValue": "https://spdx.org/rdf/terms/#checksumValue",
"ChecksumAlgorithm": "https://spdx.org/rdf/terms/#ChecksumAlgorithm"
}
],
[...]
{
"@id": "https://spdx.org/rdf/terms/#checksumAlgorithm_md5",
"@type": "ChecksumAlgorithm",
"name": "MD5",
"description": "MD5 message-digest algorithm"
},
{
"@id": "https://rocrate.s3.computational.bio.uni-giessen.de/file.txt",
"@type": "File",
"name": "file.txt",
"description": "A sample text file in the root crate.",
"encodingFormat": "text/plain",
"contentSize": 5,
"dateCreated": "2025-02-10",
"checksumAlgorithm": {"@id": "https://spdx.org/rdf/terms/#checksumAlgorithm_md5"},
"checksumValue": "a2e4822a98337283e39f7b60acf85ec9"
},
[...]
The choice of checksum algorithm was also debated, with BLAKE3 and the ISCC technical innovation being suggested as potential candidates. A remaining challenge is how to efficiently update the checksum value when the content of a data entity changes, especially for large datasets. One proposed strategy was to recalculate everything when a change was triggered; however, as this solution represents a brute force approach, it was deemed suitable only for testing purposes rather than production use. This topic remains open for future investigation, as it is crucial for maintaining consistency and synchronization across interconnected RO-Crate structures in federated environments.
Propagating Metadata Updates from Top-Level Crates to Referenced Subcrates
The question of how to propagate metadata updates from a top-level crate to its referenced subcrates was raised but not fully discussed to a final conclusion during the sessions.
Handling Property Conflicts in Nested RO-Crates
The primary applications of RO-Crate merging involve integrating distributed RO-Crates within federated systems for export purposes and enabling collaborative editing among multiple users. When independent RO-Crates are consolidated into a unified metadata document, access becomes more straightforward, yet this process alters the previously unambiguous assumptions regarding identity and provenance. Mainly the merging of separate crates introduces complications with conflicting assertions about identical entities, and relationships that span across crates. A properly merged crate must ensure that it still contains valid JSON-LD and eventually even enable that these transformations remain visible and understandable to users.
In the discussions we came up with a non-exhaustive rules to enforce during merging:
- Every described data entity must be reachable from the root via
hasPart - Ensure
@iduniqueness; merge true duplicates, or mint new identifiers and record original ids as provenance - Preserve differing property values by representing the property as an array and marking each value with its source
- tbc
Results
Extensions to ro-crate-py
Extended the library with a class Subcrate extending the Dataset class. This additional implementation allows things like:
main_crate = ROCrate("/tmp/ro-crate-dir")
subcrate = main_crate.get("subcrate")
subfile = subcrate.get("subfile.txt")
# or
subfile = subcrate["hasPart"][0]
# or
entities = subcrate.get_entities()
The implementation is such that the subcrate is only loaded when accessing some of its attribute, to avoid potentially loading large amount of metadata, as one purpose of the subcrate is also to reduce the amount of information in the main crate.
🔗 Link to the pull request with the feature implementations
Extensions to ro-crate-rs
We added an interactive CLI mode for RO-Crate (attached and detached) exploration and traversal. This integrates the following commands:
Commands:
load <url|path> Load RO-Crate from URL or .zip file
ls List hasPart of current dataset/folder
ls -a List all entities (data + contextual)
get <@id> Pretty-print JSON for entity
cd <id> Enter subcrate or folder
cd .. Return to parent
cd / Return to root crate
pwd Print current path
help Show this message
Supported formats:
- .zip archives containing ro-crate-metadata.json
- Direct URLs to RO-Crate archives
- DOIs resolving to Zenodo/similar repositories
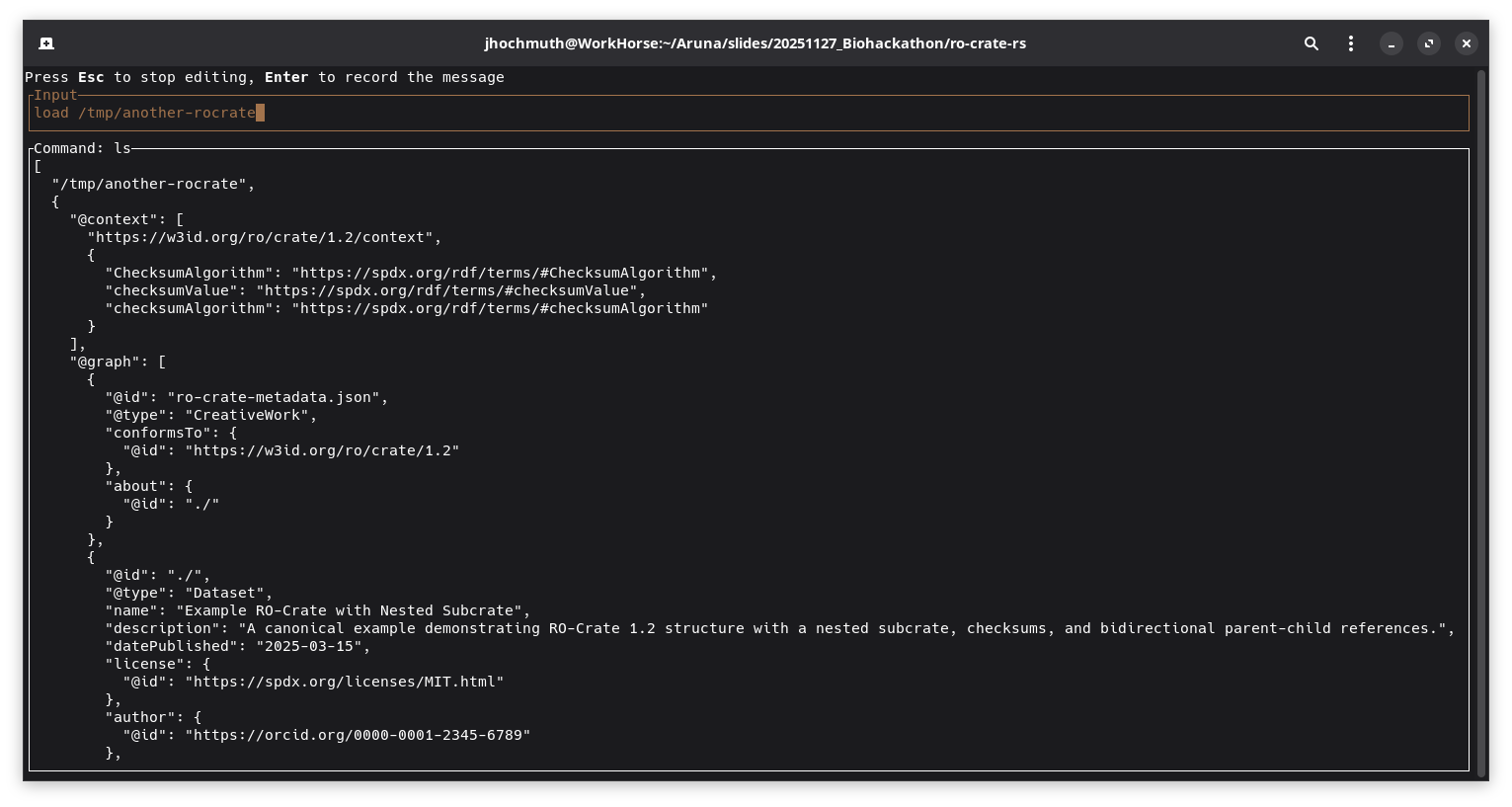
Screenshot which shows the complete loaded top-level RO-Crate metadata file
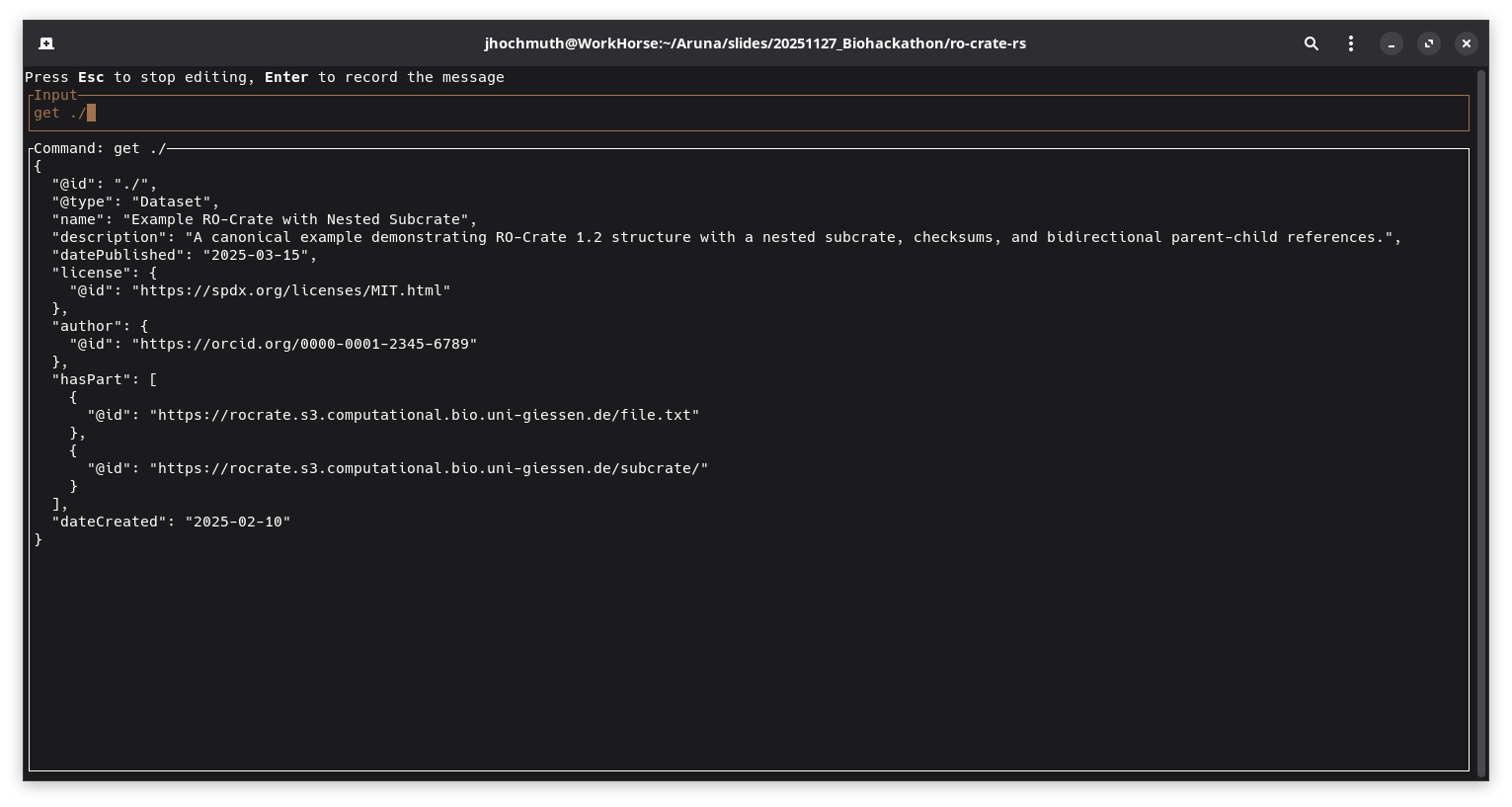
Screenshot which displays a specific entity after executing the get ./ command
During the development process, discussions were held with one of the official RO-Crate specification maintainers regarding the potential transfer of the library into the official ResearchObject Github organization. This conversation explored the benefits of housing the library within the established organizational structure, which would provide greater visibility within the RO-Crate community and signal its alignment with official specification standards. Future discussions with the current maintainer will focus on establishing joint development and shared maintenance of the library.
RO-Crate indexing for easier exploration
The rocrate-indexer implements a thin webserver that provides the following endpoints:
[POST] /crates/url Add an RO-Crate from a URL to the index
[POST] /crates/upload Add an RO- to the index by uploading a file (zip archive or ro-crate-metadata.json)
[GET] /crates List information of all indexed RO-Crates
[GET] /crates/{crate-id} Get full metadata of the RO-Crate associated with the specific id
[GET] /crates/{crate-id}/info Get shortened information (name, description, ancestry path) of the RO-Crate associated with the specific id
[GET] /search Search for entities matching a query
[DELETE] /crates/{crate-id} Delete RO-Crate associated with the specific id from the index
The tool integrates a fulltext search index over all included entities of an RO-Crate metadata file. The search functionality can be performed either as a fuzzy search without a cleanly defined scope or as an exact search on specific fields of entities.
Examples for search queries:
-
- Full text search
- “e.coli”
-
- Search by type
- “entity_type:Person”
-
- Search by nested property
- “author.name:Smith”
-
- Boolean query
- “name:Test AND entity_type:Dataset”
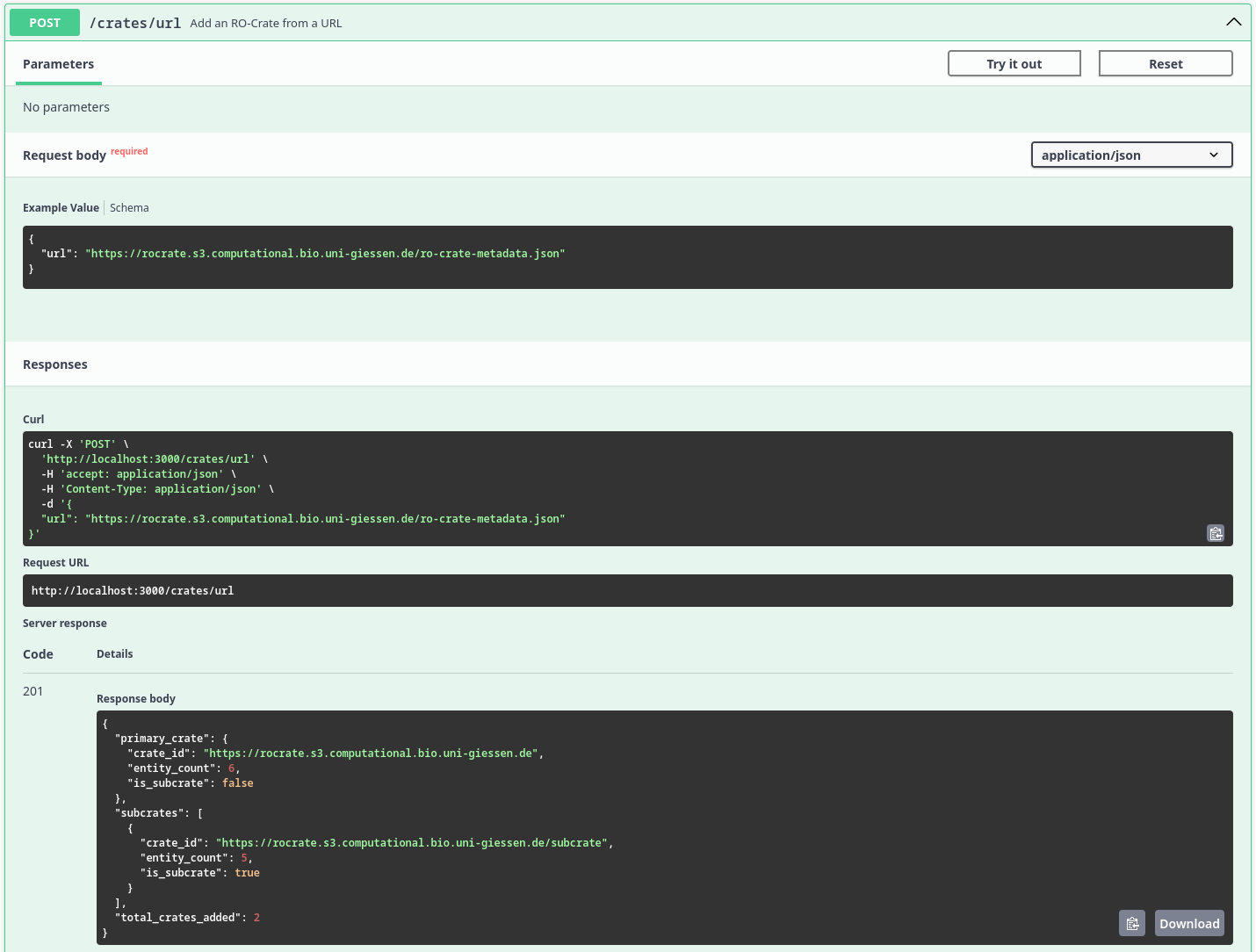
Screenshot which displays the result of the /crates/url endpoint after recursively ingesting the RO-Crate metadata files
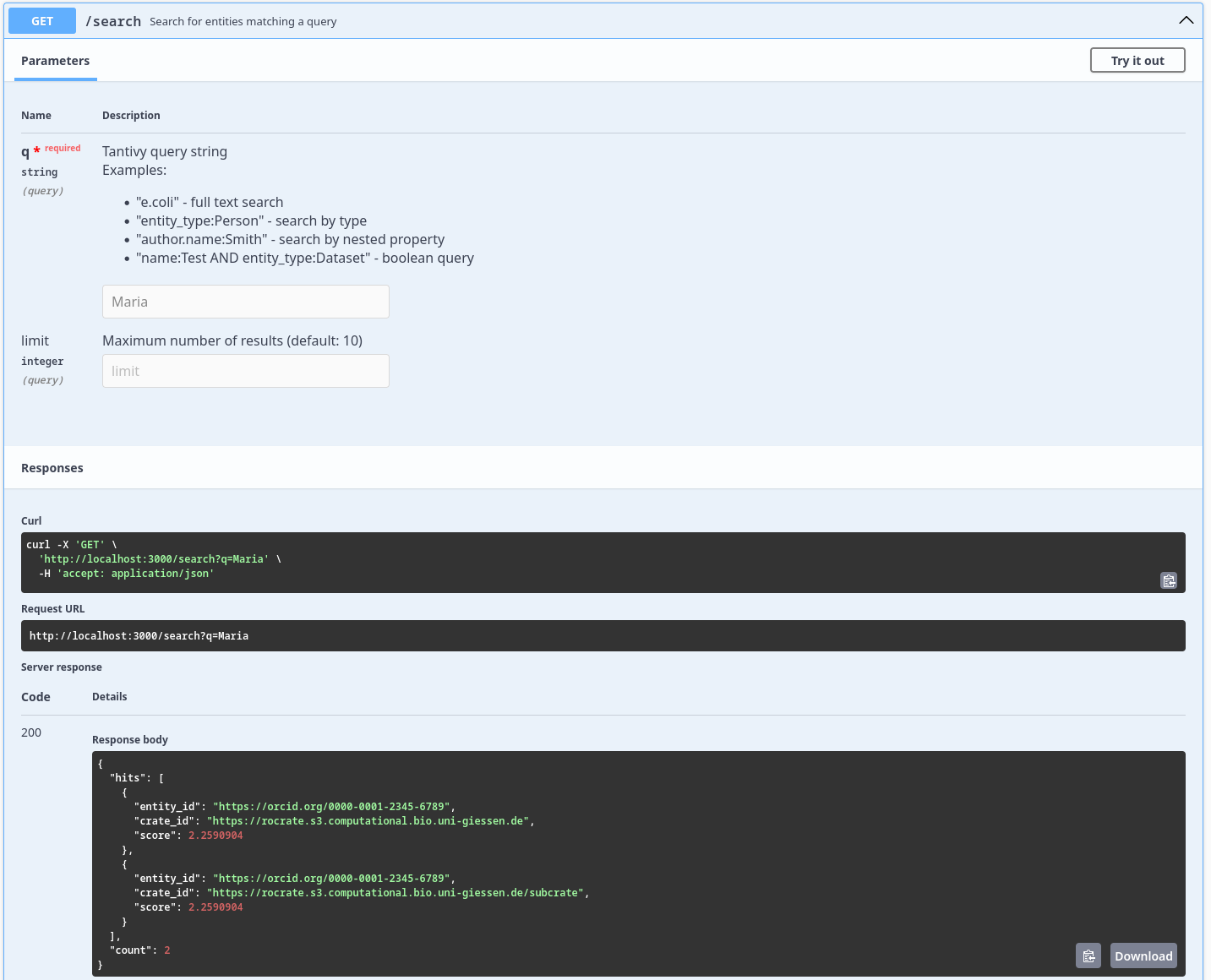
Screenshot which displays the result of the /search endpoint after searching for the query Maria
Demonstrator
The RO-Crate Explorer is a lightweight “Minimum Viable Product” (MVP) web application designed to demonstrate the parsing, visualization, and traversal of Research Object Crates (RO-Crates). It provides a user-friendly interface to navigate the complex graph structures of metadata, supporting both detached (remote) and local nested crate structures.
Core Capabilities
1. Universal Input Methods
The application serves as a flexible entry point for RO-Crate data, supporting three distinct loading mechanisms:
- Remote URL: Users can input a URL pointing to a remote crate or a specific
ro-crate-metadata.jsonfile. This allows for the exploration of detached crates hosted on external servers. - ZIP Archive: Users can upload a zipped RO-Crate. The application utilizes
JSZipto unpack the archive in memory and locate the metadata file automatically. - JSON Metadata: Users can upload a raw
ro-crate-metadata.jsonfile directly for immediate parsing.
2. File Tree & Graph Traversal
Once a crate is loaded, the application constructs a navigable representation of the data:
- File Tree: The sidebar displays a hierarchical tree view of the crate’s content (Datasets and Files). This is computed by traversing the graph starting from the Root Dataset, handling parent-child relationships via the
hasPartproperty. - Nested Crate Support: The application is capable of detecting links to other RO-Crates. If an entity links to another
ro-crate-metadata.json, the interface provides a dedicated “Explore Subcrate” button. Clicking this pushes the current state to a history stack and loads the new crate context, allowing users to “dive” deep into nested structures.
3. Context & Filtering
Beyond the directory structure, RO-Crates contain rich contextual entities (e.g., Person, Organization, CreativeWork).
- Context Filter: The sidebar acts as a filter, grouping all non-file entities by their
@type. - Graph Access: This allows users to quickly locate specific metadata entities that do not appear in the physical file hierarchy but are crucial to the graph.
4. Entity Inspection & Preview
Clicking any item in the tree or context menu opens the Entity Viewer:
- Property Preview: Displays all JSON-LD properties associated with the entity (e.g.,
author,datePublished,description). - Link Navigation: Properties that link to other entities (by
@id) are clickable, allowing the user to jump between connected nodes in the graph. - Raw Data: A “Raw JSON” option is available to inspect the underlying JSON-LD source for debugging purposes.
Process Flow
The following diagram illustrates how the Demonstrator processes user input and navigates the crate graph.
graph TD
A[User Entry] -->|Upload .zip/.json| B(File Parser)
A -->|Enter URL| C(Fetch API)
B --> D{Parse JSON-LD}
C --> D
D --> E[RO-Crate Object Model]
E --> F[Generate File Tree]
E --> G[Group Entities by Type]
F --> H[UI: Sidebar Navigation]
G --> H
H -->|Select Entity| I[UI: Entity Detail View]
I -->|Click Link| I
I -->|Click Subcrate Link| J[Push to History Stack]
J --> C
Interface Overview
Dashboard State
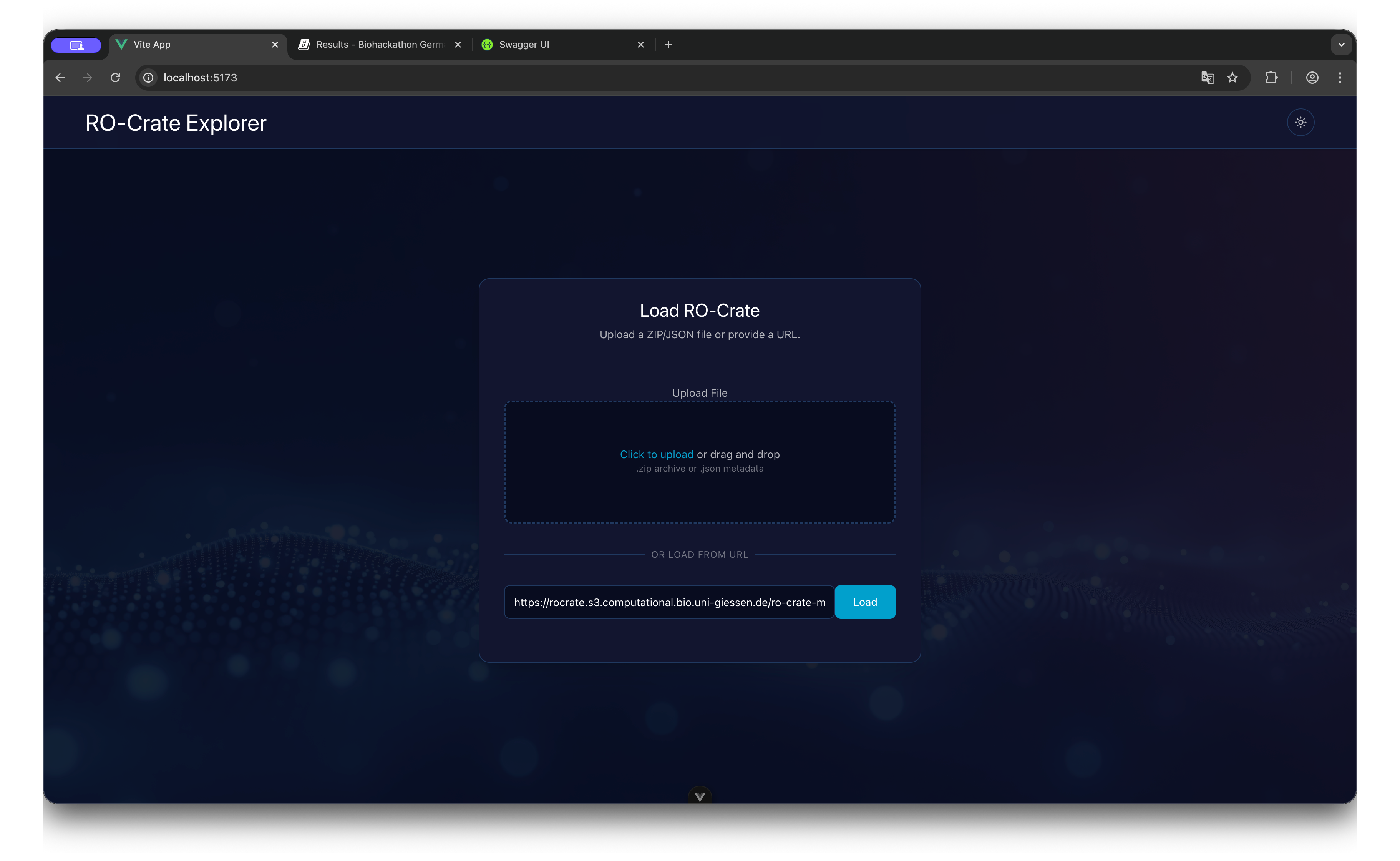
The entry screen allows for dragging and dropping files or pasting a remote URL to initialize the session.
Explorer State
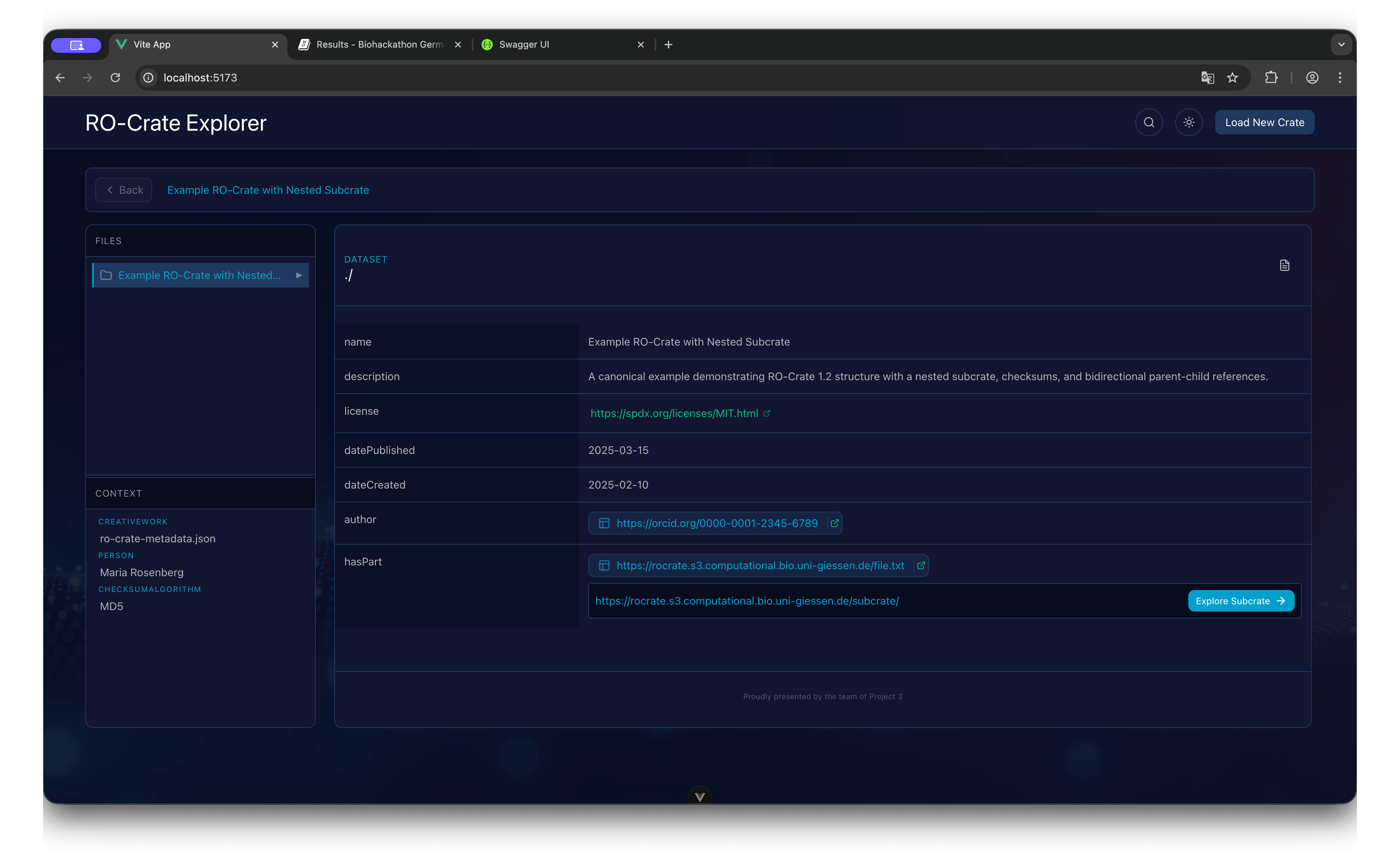
- Header: Displays the current Crate name and Breadcrumb navigation history.
- Sidebar (Files): A collapsible tree structure representing the physical file organization.
- Sidebar (Context): A list of non-file entities (e.g., ContextEntity, Person) grouped by type.
- Main Content: The detailed view of the currently selected entity, showing key-value pairs of metadata.
Nested Navigation
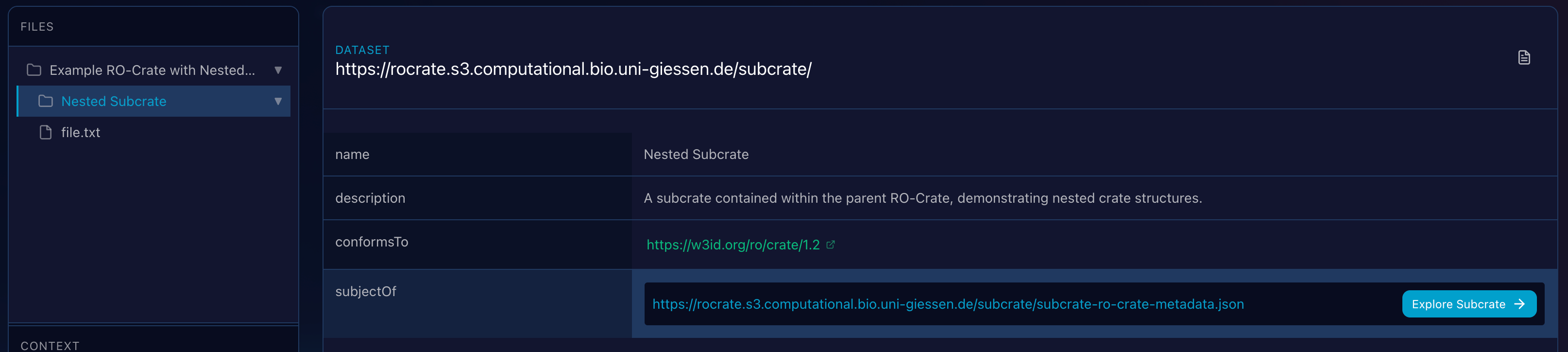
When the viewer encounters a property linking to another Metadata file, it renders an action button to seamlessly transition the application context to that sub-crate.
Full-Text Search in RO-Crate Explorer
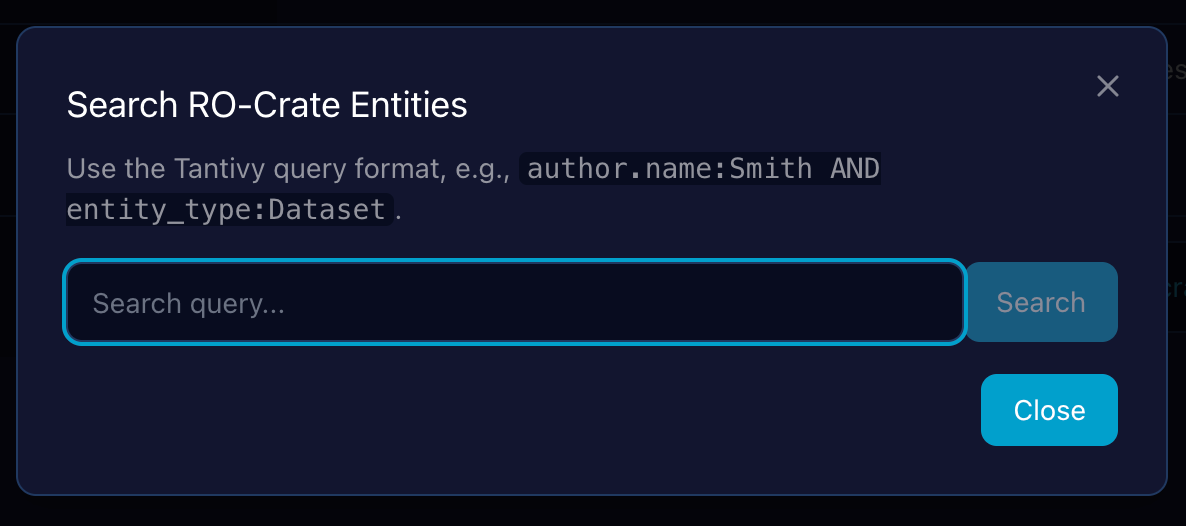
In addition to traversal, the Demonstrator provides a full-text search capability that indexes all metadata across the loaded crate, allowing users to quickly find entities by matching terms in their properties (e.g., name, description, author).
Linked Entity Inspection as an Overlay
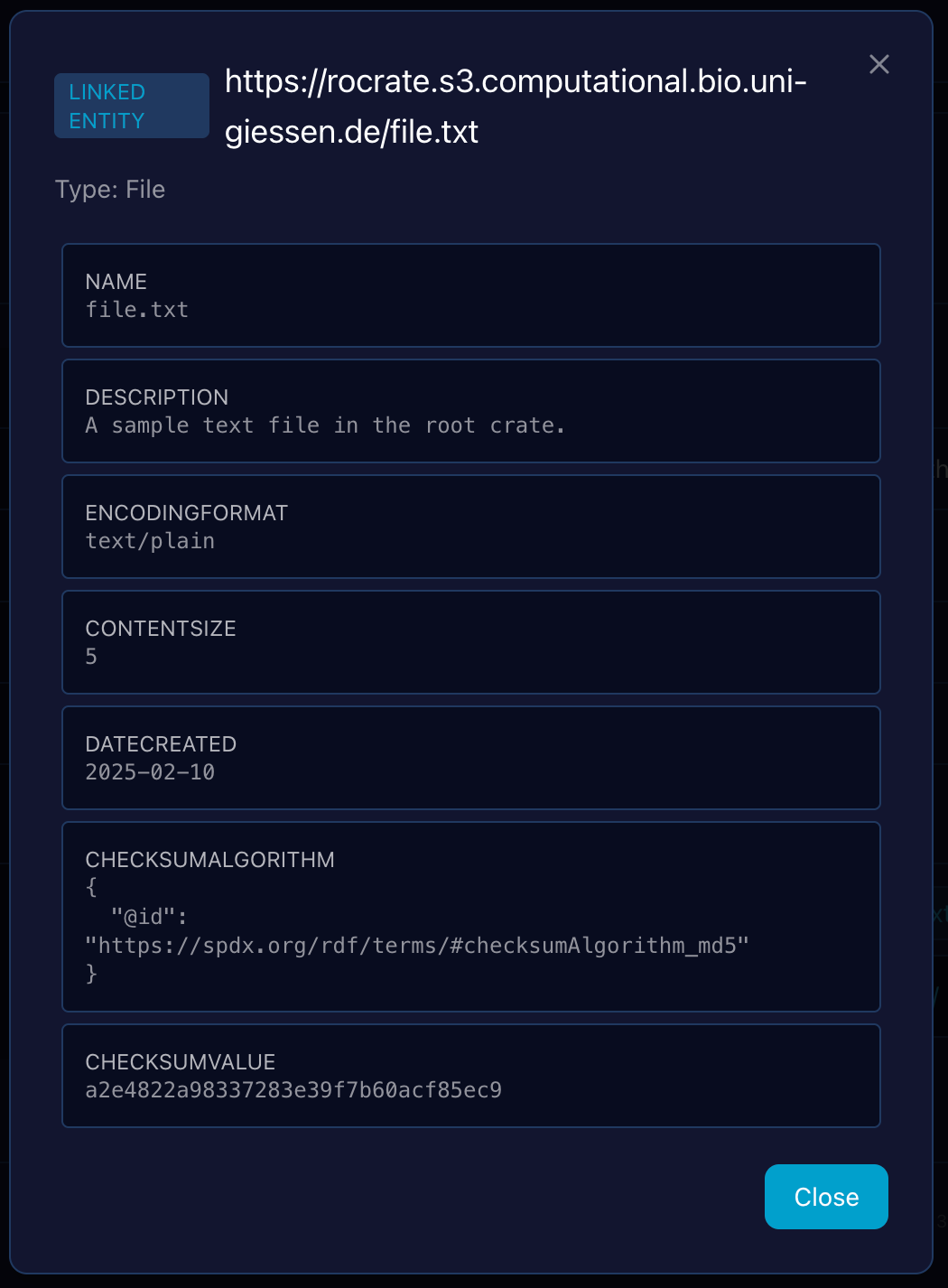
To maintain context during exploration, clicking on an entity link (an @id reference) within the main Entity Detail View triggers an overlay window rather than navigating away. This allows users to quickly inspect the metadata of a linked entity (e.g., an author or a datePublished entity) without losing the context of the parent entity they were initially viewing.
Raw JSON-LD Source Overlay and Export Actions
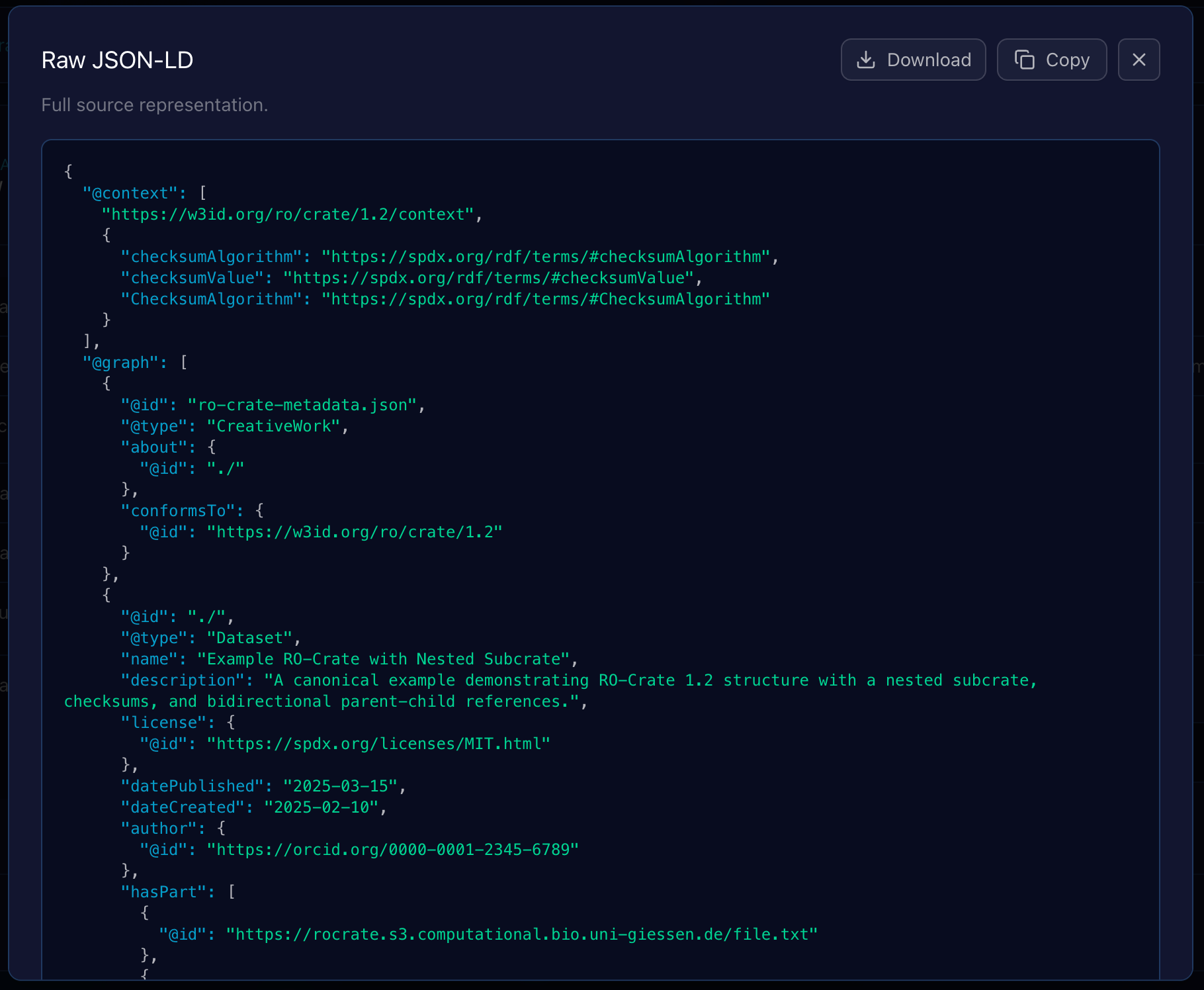
Clicking the “Raw JSON” option within the Entity Viewer opens a modal overlay that displays the entity’s complete, unparsed JSON-LD source code. This feature is crucial for debugging, schema validation, and verifying the exact structure of the data as it appears in the ro-crate-metadata.json file.
-
- Lightweight Syntax Highlighting
- The raw JSON-LD is rendered with basic syntax highlighting to significantly improve readability and help users quickly parse the structure of keys, strings, and values.
-
- Debug and Validation
- This view offers an immediate, faithful representation of the data, allowing users to confirm the properties and values being loaded by the application’s internal object model.
To facilitate working with this raw data outside the application, the overlay is equipped with two export actions:
1. Copy to Clipboard
A prominent “Copy” button is included within the overlay.
Functionality: Clicking this button instantly copies the entire, raw JSON-LD source text from the overlay to the user’s clipboard, ready to be pasted into a text editor, debugger, or other tool.
2. Download Entity Data
A “Download” button is provided to save the crate’s metadata locally.
Format: The downloaded file contains the complete, raw entity metadata as a standard JSON-LD fragment.
These export functions ensure that developers and users can easily access and manipulate the exact metadata harvested by the RO-Crate Explorer.